All content on this site is intended for healthcare professionals only. By acknowledging this message and accessing the information on this website you are confirming that you are a healthcare professional. If you are a patient or carer, please visit the MDS Alliance.
The MDS Hub website uses a third-party service provided by Google that dynamically translates web content. Translations are machine generated, so may not be an exact or complete translation, and the MDS Hub cannot guarantee the accuracy of translated content. The MDS Hub and its employees will not be liable for any direct, indirect, or consequential damages (even if foreseeable) resulting from use of the Google Translate feature. For further support with Google Translate, visit Google Translate Help.
Now you can support HCPs in making informed decisions for their patients
Your contribution helps us continuously deliver expertly curated content to HCPs worldwide. You will also have the opportunity to make a content suggestion for consideration and receive updates on the impact contributions are making to our content.
Find out more
Create an account and access these new features:
Bookmark content to read later
Select your specific areas of interest
View MDS content recommended for you
Targeted next-generation sequencing: A comprehensive genetic profiling tool to simultaneously detect copy number variants and somatic mutations in myeloid malignancies
Next-generation sequencing (NGS) is a high throughput analytical tool used for diagnosis of hereditary disorders such as myeloid malignancies. Copy number variants (CNVs) and genetic aberrations are used as a prognostic biomarker in myeloid malignancies. A targeted NGS assay is widely employed to identify targeted mutations, such as point mutations/single-nucleotide variants, insertions, and deletions. However, the potential use of the targeted NGS assay to detect common CNVs in myeloid malignancies compared with the traditional fluorescence in situ hybridization (FISH) panel, and conventional chromosome analysis (CCA) is still not completely determined.
In a recent proof-of-principle study, published in The Journal of Molecular Diagnostic, Liqun Jiang and colleagues1 compare the detection of myelodysplastic syndrome (MDS) CNVs by a targeted NGS with routine clinical methods (MDS FISH, and CCA). The study aimed to explore the possibility of using targeted NGS to identify somatic mutations and CNVs at the same time.
Methods
Samples of bone marrow or blood from 406 patients were analyzed using MDS FISH, targeted NGS, and CCA, and only 91 specimens with an abnormal MDS FISH panel were selected for the study. However, 315 normal samples were further characterized using CCA and targeted NGS. Figure 1 describes an outline of the methods employed in the study.
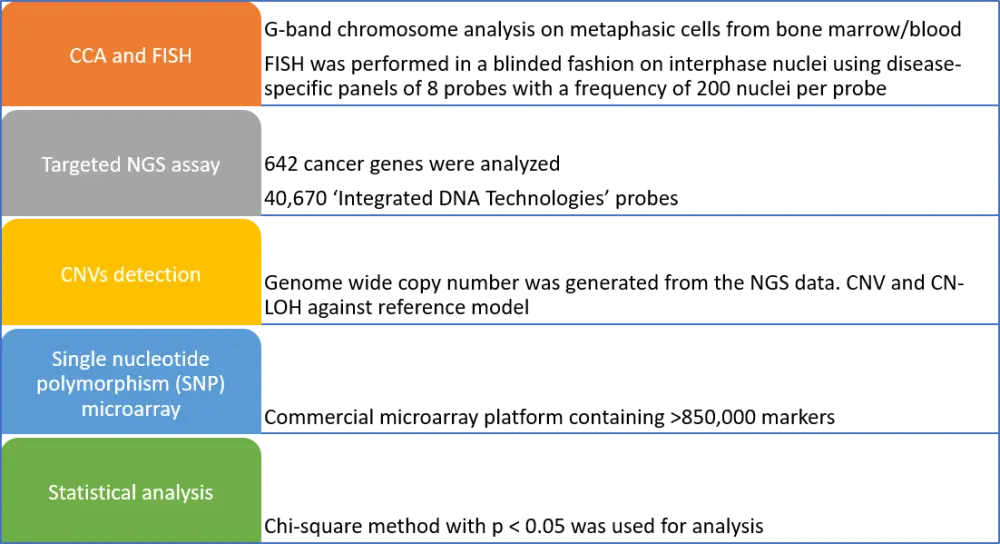
CCA, conventional chromosome analysis; CN-LOH, copy neutral-loss of heterozygosity; CNV, copy number variant; FISH, fluorescent in situ hybridization; NGS, next-generation sequencing.
*Adapted from L. Jiang et al.1
Results
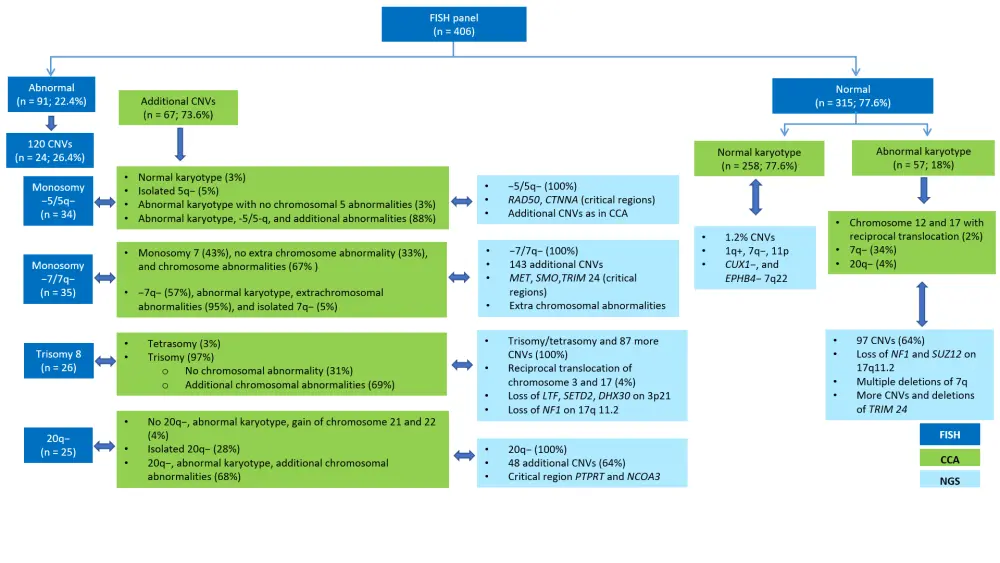
A succinct summary of the results is presented in Figure 2, and key outcomes are as follows:
- Among 315 normal cases as per FISH panel, 57 had abnormal karyotype. In 258 patients with normal karyotypes, three cases were identified to carry CNVs as identified by targeted NGS.
- In total, 91 abnormal cases were detected by CCA and MDS FISH, and the targeted NGS was in complete agreement to the outcomes of both conventional methods.
- Targeted NGS also identified missed mutations in a number of cases, either due to the fact that these deletions were cryptic and were beyond the limits of CCA, or not covered by the FISH probes. Two cases had 5q deletions, one case had 20q deletions, and one case had a cryptic deletion of 7q22 in CUX1 and EPHB4 genes. Three-quarters of patients were reported to have deletions in the CUX1 and EPHB4 genes along with −7/7q deletions.
- Patients with 7q abnormalities also had deletions in the MET, SMO, and more commonly TRIM24, genes; it is advised that evaluation of neoplastic transformation in patients with 7q deletions, and regions around the TRIM24 gene and co-occurring gene mutations may provide more information about pathogenesis in myeloid malignancies.
- Copy neutral loss of heterozygosity (CN-LOH), which is associated with a higher risk of disease recurrence, could not be picked by the two conventional methods. For instance, the targeted NGS assay identified one case with a CN-LOH in chromosome 13 with an internal tandem duplication (ITD) of the FLT3 gene on chromosome 13q, and a variant allele fraction (VAF) of approximately 79%. This suggests duplication of oncogenic mutations and loss of normal alleles of genes.
Limitations
- In the study,1 the lower detection limit of the NGS assay was 5%, 3.5%, 4%, and 4.5% for monosomy 5/5q deletions, monosomy 7/7q deletions, trisomy 8, and 20q deletions, respectively. Detection limit depends upon a number of factors such as proportion of abnormal cells, proportion of the subclonal population containing the CNVs, genome ploidy, and the size of the CNVs.
- The proposed targeted NGS assay was unable to identify balanced structural abnormalities, such as balanced reciprocal translocations, and inversions; this issue can be addressed in the future with the addition of probes to capture these abnormalities in myeloid malignancies.
- The targeted NGS assay provides an estimated percentage of abnormal cells (VAF) instead of quantitative percentage of abnormal cells as provided by FISH.
Conclusion
Targeted NGS is a highly sensitive assay that provides a single platform to detect CNVs, relevant somatic mutations, and CN-LOH in myeloid malignancies in a single experiment. The assay offers a quick and prompt way of identification of critical regions, and can detect mutations missed by CCA or the MDS FISH panel.
References
Please indicate your level of agreement with the following statements:
The content was clear and easy to understand
The content addressed the learning objectives
The content was relevant to my practice
I will change my clinical practice as a result of this content

